Phytoplankton species classifier
Classify phytoplankton images among 60 classes.
Model | Trainable | Inference | Pre-trained
Published by
DEEP-Hybrid-DataCloud Consortium
Created:
- Updated:
Model Description
Citizen science has become a powerful force for scientific inquiry, providing researchers with access to a vast array of data points while connecting non scientists to the real process of science. This citizen-researcher relationship creates a very interesting synergy, allowing for the creation, execution, and analysis of research projects. With this in mind, a Convolutional Neural Network has been trained to identify phytoplankton in collaboration with the Vlaams Instituut voor de Zee.
This Docker container contains a trained Convolutional Neural network optimized for phytoplankton identification using images. The architecture used is an Xception [1] network using Keras on top of Tensorflow.
The PREDICT method expects an RGB image as input (or the url of an RGB image) and will return a JSON with the top 5 predictions.
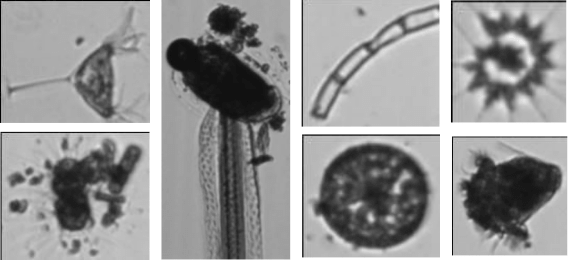
As training dataset we have used a collection of images from the Vlaams Instituut voor de Zee which consists of around 650K images from 60 classes of phytoplankton.
This service is based in the Image Classification with Tensorflow model.
References
[1]: Chollet, Francois. Xception: Deep learning with depthwise separable convolutions arXiv preprint (2017): 1610-02357.
Test this module
You can test and execute this module in various ways.
Excecute locally on your computer using Docker
You can run this module directly on your computer, assuming that you have Docker installed, by following these steps:
$ docker pull deephdc/deep-oc-phytoplankton-classification-tf
$ docker run -ti -p 5000:5000 deephdc/deep-oc-phytoplankton-classification-tfExecute on your computer using udocker
If you do not have Docker available or you do not want to install it, you can use udocker within a Python virtualenv:
$ virtualenv udocker
$ source udocker/bin/activate
(udocker) $ pip install udocker
(udocker) $ udocker pull deephdc/deep-oc-phytoplankton-classification-tf
(udocker) $ udocker create deephdc/deep-oc-phytoplankton-classification-tf
(udocker) $ udocker run -p 5000:5000 deephdc/deep-oc-phytoplankton-classification-tf
In either case, once the module is running, point your browser to
http://127.0.0.1:5000/ and you will see the API
documentation, where you can test the module functionality, as well as
perform other actions (such as training).
For more information, refer to the user documentation.
Train this module
You can train this model using the DEEP framework. In order to execute this module in our pilot e-Infrastructure you would need to be registered in the DEEP IAM.
Once you are registedered, you can go to our training dashboard to configure and train it.
For more information, refer to the user documentation.